The Journey of Data: Lessons Learned in Modeling Kinase Affinity, Selectivity, and Resistance [Article v1.0]
DOI:
https://doi.org/10.33011/livecoms.6.1.3875Keywords:
Structure-based ML, ML drug discovery, Kinases, Binding affinity predictions, Drug discovery framework, Protein-ligand binding, Data harmonizationAbstract
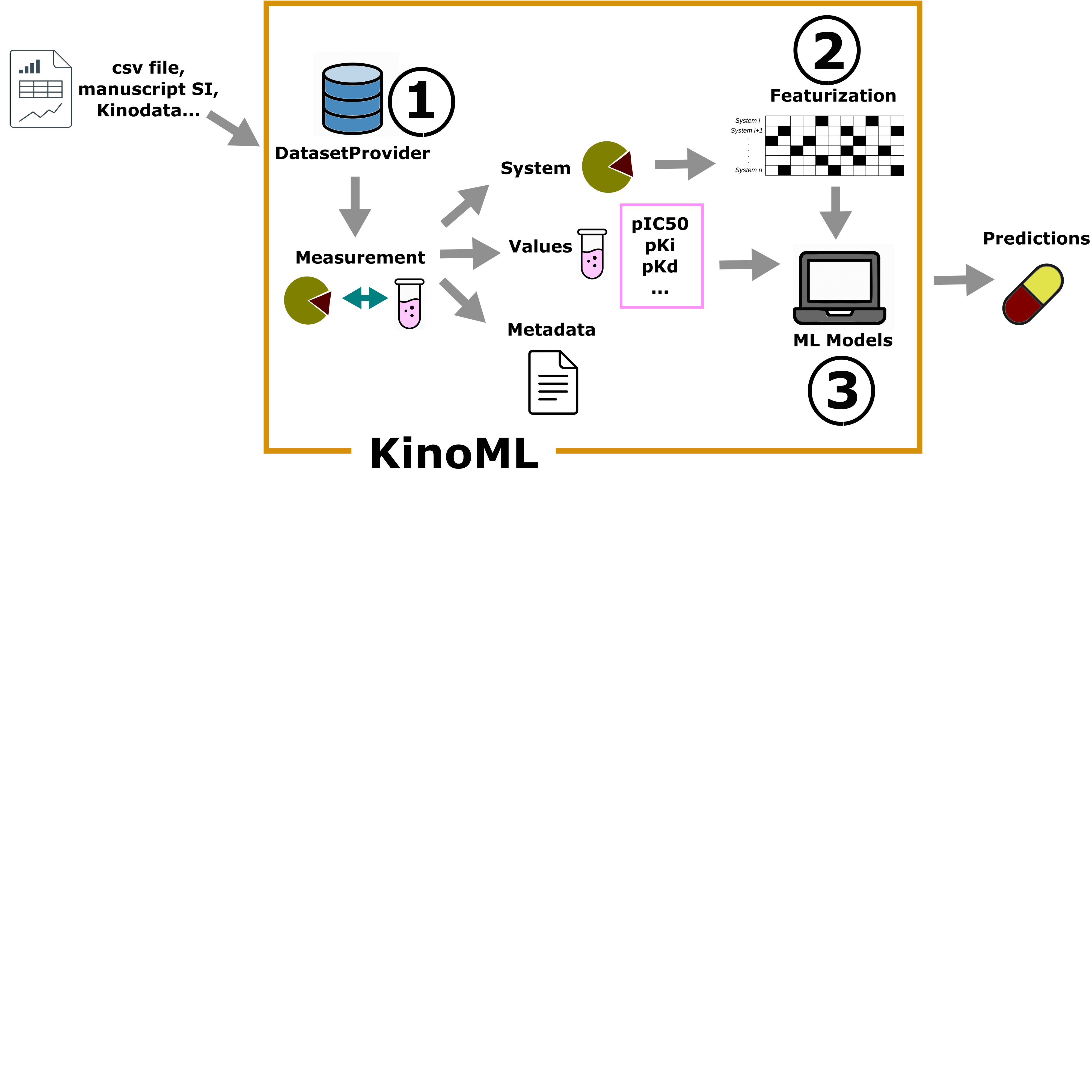
Recent advances in machine learning (ML) are reshaping drug discovery. Structure-based ML methods use physically-inspired models to predict binding affinities from protein:ligand complexes. These methods promise to enable the integration of data for many related targets, which addresses issues related to data scarcity for single targets and could enable generalizable predictions for a broad range of targets, including mutations. In this work, we report our experiences in building KinoML, a novel framework for ML in target-based small molecule drug discovery with an emphasis on structure-enabled methods. KinoML focuses currently on kinases as the relative structural conservation of this protein superfamily, particularly in the kinase domain, means it is possible to leverage data from the entire superfamily to make structure-informed predictions about binding affinities, selectivities, and drug resistance. Some key lessons learned in building KinoML include the importance of reproducible data collection and deposition, the harmonization of molecular data and featurization, and the selection of the right data format to ensure reusability and reproducibility of ML models. As a result, KinoML allows users to easily achieve three tasks: accessing and curating molecular data; featurizing this data with representations suitable for ML applications; and running reproducible ML experiments that require access to ligand, protein, and assay information to predict ligand affinity. Despite KinoML focusing on kinases, this framework can be applied to other proteins. The lessons reported here can help guide the development of platforms for structure-enabled ML in other areas of drug discovery.
Downloads
Published
How to Cite
License
Copyright (c) 2025 Raquel López-Ríos de Castro, Jaime Rodríguez-Guerra, David Schaller, Talia B. Kimber, Corey Taylor, Jessica B. White, Michael Backenköhler, Joschka Groß, Alexander M. Payne, Benjamin Kaminow, Ivan Pulido, Sukrit Singh, Paula Linh Kramer, Guillermo Pérez-Hernández, Andrea Volkamer, John D. Chodera

This work is licensed under a Creative Commons Attribution 4.0 International License.

