Structure-Based Experimental Datasets for Benchmarking Protein Simulation Force Fields [Article v1.0]
DOI:
https://doi.org/10.33011/livecoms.6.1.3871Keywords:
Molecular simulation, Force field, X-ray crystallography, NMR spectroscopyAbstract
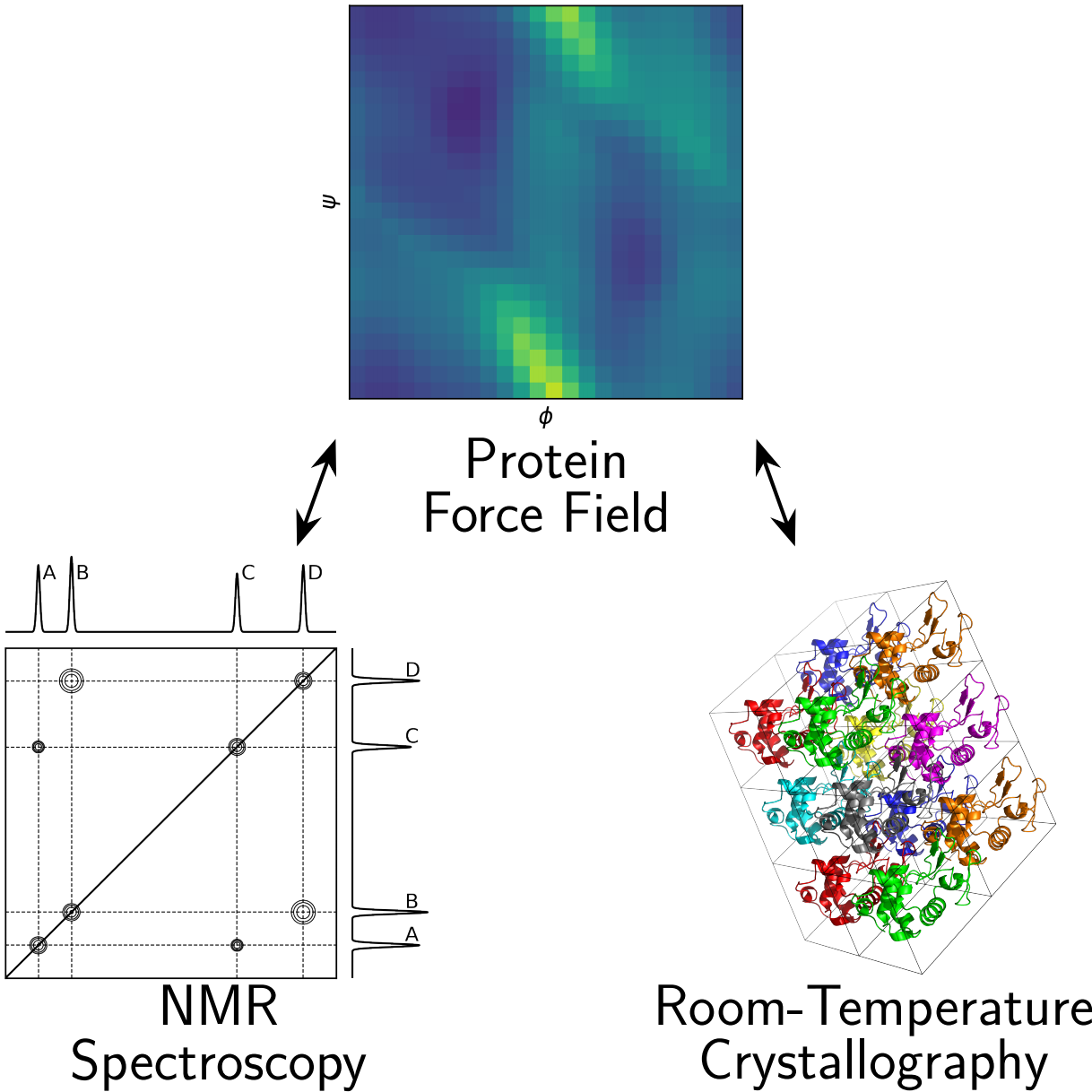
This review article provides an overview of structurally oriented experimental datasets that can be used to benchmark protein force fields, focusing on data generated by nuclear magnetic resonance (NMR) spectroscopy and room temperature (RT) protein crystallography. We discuss what the observables are, what they tell us about structure and dynamics, what makes them useful for assessing force field accuracy, and how they can be connected to molecular dynamics simulations carried out using the force field one wishes to benchmark. We also touch on best practices for setup and analysis of benchmark simulations. We hope this article will be particularly useful to computational researchers and trainees who develop, benchmark, or use protein force fields or machine learning models that generate protein ensembles.
Downloads
Published
How to Cite
License
Copyright (c) 2025 Chapin E. Cavender, David A. Case, Julian C.-H. Chen, Lillian T. Chong, Daniel A. Keedy, Kresten Lindorff-Larsen, David L. Mobley, O. H. Samuli Ollila, Chris Oostenbrink, Paul J. Robustelli, Vincent A. Voelz, Michael E. Wall, David C. Wych, Michael K. Gilson

This work is licensed under a Creative Commons Attribution 4.0 International License.

