Quantifying Spatially Resolved Hydration Thermodynamics Using Grid Inhomogeneous Solvation Theory [v1.0]
DOI:
https://doi.org/10.33011/livecoms.6.1.3059Keywords:
Grid Inhomogeneous Solvation Theory, GIST, Solvation Free Energy, Tutorial, Protein Ligand BindingAbstract
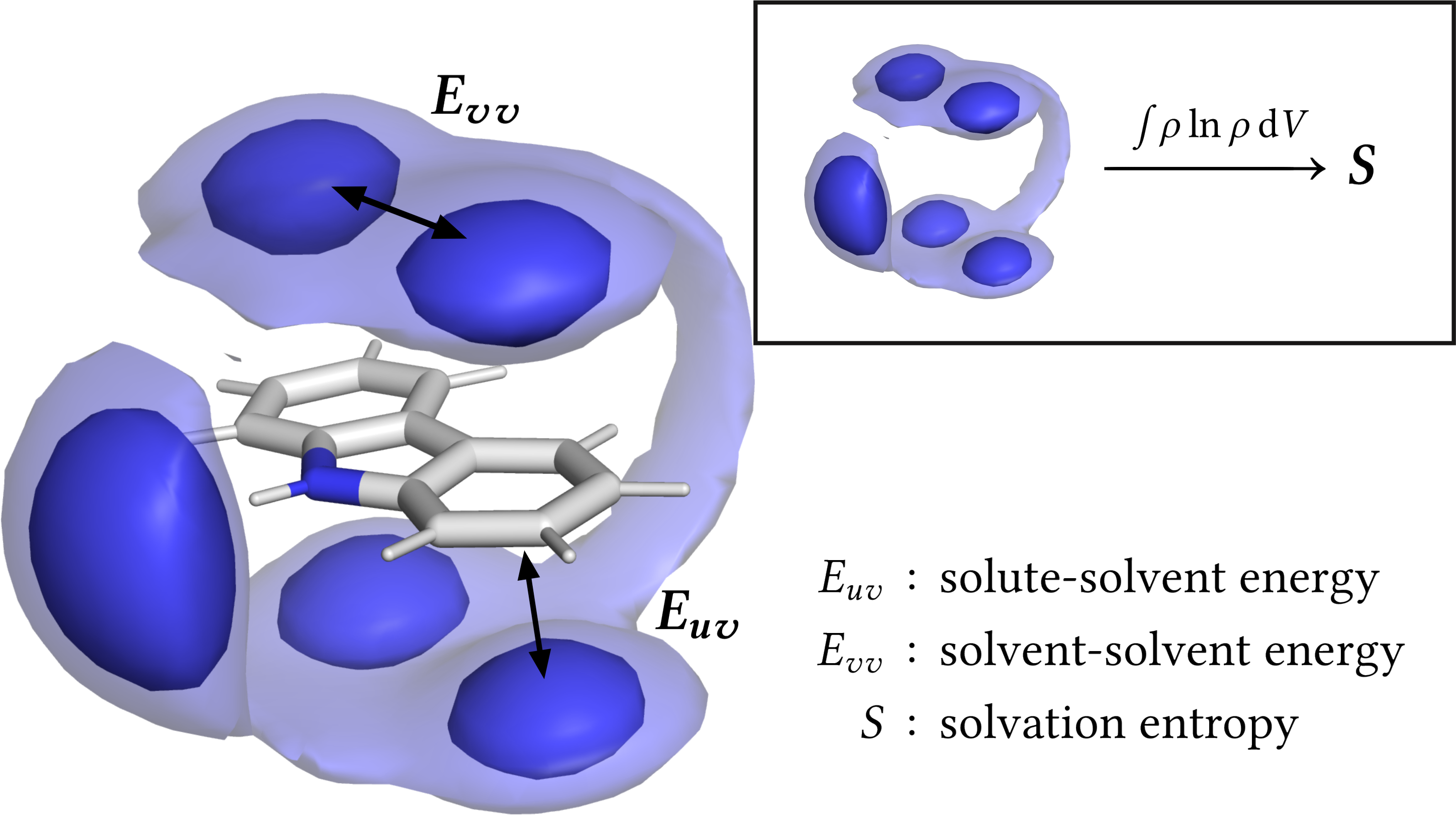
Grid Inhomogeneous Solvation Theory (GIST) is a method to compute the free energy of solvation of a solute molecule on a three-dimensional grid based on sampling from molecular dynamics (MD) simulations. The high spatial resolution of the GIST output, as well as the decomposition into energy and entropy contributions, allow for highly detailed analyses of solvation around both proteins and small molecules. However, this versatility also comes with a significant entry barrier for new users.
In this tutorial, we aim to guide the reader through the most common steps involved in a GIST analysis using the streptavidin-biotin complex as a demonstrative system. To this end, Jupyter notebooks and a Python package (gisttools) are provided to simplify the analysis. Furthermore, we discuss the theory of GIST with a focus on practical aspects. We highlight potential pitfalls and provide strategies to avoid technical difficulties. This tutorial assumes familiarity with molecular dynamics simulations and the AmberTools package.
Downloads
Published
How to Cite
License
Copyright (c) 2025 Valentin J. Egger-Hoerschinger, Franz Waibl, Vjay Molino, Helmut Carter, Monica L. Fernández-Quintero, Steven Ramsey, Daniel R. Roe, Klaus R. Liedl, Michael K Gilson, Tom Kurtzman

This work is licensed under a Creative Commons Attribution 4.0 International License.

