Lessons Learned from the Calculation of One-Dimensional Potentials of Mean Force [Article v1.0]
DOI:
https://doi.org/10.33011/livecoms.1.2.11073Keywords:
potential of mean force, binding free energy, artifactsAbstract
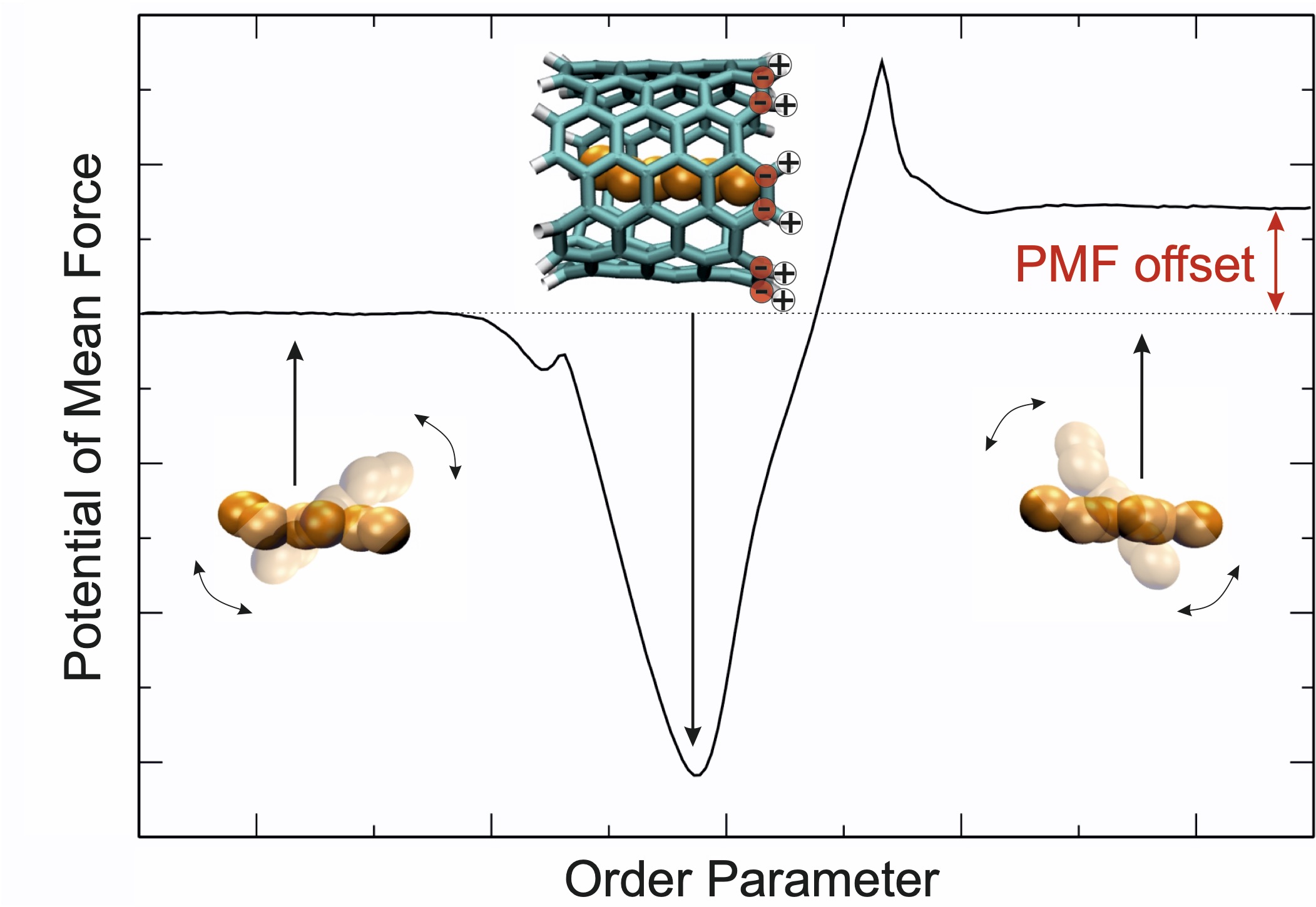
The origins of different computational artifacts that may occur in the calculation of one-dimensional potentials of mean force (PMF) via umbrella sampling molecular dynamics simulations and manifest as free energy offset between bulk solvent regions are investigated. By systematic studies, three distinct causes are elucidated: (i) an unfortunate choice of reference points for the umbrella distance restraint; (ii) a misfit in probability distributions between bound and unbound umbrella windows in case of multiple binding modes; (iii) artifacts introduced by the free energy estimator. Starting with a fully symmetric model system consisting of methane binding to a cylindrical host, complexity is increased through the introduction of dipolar interactions between the host and the solvent, the host and the guest molecule or between all involved species, respectively. The manifestation of artifacts is illustrated and their origin and prevention is discussed. Finally, the consequences for the calculation of standard binding free enthalpies is illustrated using the complexation of primary alcohols with alpha-cyclodextrin as an example.